If you’ve followed any of my work for the past couple of years, you’ll know I used to work at Kuzu, where we were building an embedded graph database. My work at Kuzu was all about helping users figure out how to transform and query their data as a graph, experimenting with how to define graph data models and relationships for different domains, how to bring the power of Cypher to developers who didn’t want to spin up a server or pay for a proprietary tool just to run traversal queries.
In September 2025, Kuzu got archived, and the codebase went quiet. But the ideas were still relevant. The graph community still highly valued a permissively licensed, embedded graph storage and query engine that supported Cypher.
That’s where LadybugDB comes in. It’s a fork of Kuzu, maintained by the team at Ladybug Memory ⤴, but with a slightly different goal. Kuzu was built primarily as a standalone, embedded graph database. Ladybug wants to be a graph lakehouse — interoperating with DuckDB’s storage layer, reading and writing Arrow/Parquet, connecting to object stores. The same columnar foundation, but opened up to the broader ecosystem.
Over the past few months, I’ve been in close discussions with Arun Sharma ⤴, lead maintainer of LadybugDB. Having explored what his vision looks like in practice, I wanted to share a small project that demonstrates at least a part of the Ladybug’s vision: running fast graph algorithms natively on top of columnar graph storage.
Towards the end of this post, we’ll lead into the bigger picture and what this all means for folks working with graphs.
Dataset: Who Mentored the Most Nobel Laureates?#
The dataset we’ll be working with (see the repo ⤴ if you want to follow along) is a mentorship network of Nobel laureates, containing thousands of records that trace the academic lineages of prize winners across Physics, Chemistry, and Medicine.
It turns out that research has shown Nobel Prize committees don’t just reward isolated genius. They reward people who are already in the right networks, having received mentorship from the right teachers, who were part of the just the right intellectual institutions. This makes sense when you learn from the data that Niels Bohr mentored ten Nobel laureates in Physics. Or that J.J. Thomson, discoverer of the electron, mentored eleven? These aren’t just names on a list — they’re nodes in a network of knowledge transmission that spans more than a century.
The data comes in JSON format, structured as parent-child relationships. Each record has a “child” (the mentee) and one or two “parents” (the mentors). Some of those children are laureates with prize categories and years attached. Others are scholars — people who never won a Nobel but who mentored someone who did, or who mentored someone who mentored someone who did.
This data is naturally analyzed as a graph. You could flatten it into a table and run SQL queries with traditional joins, but you’d lose its inherent structure entirely. Questions like “who are all the academic descendants of Max Planck?” or “which scholars sit at the crossroads of multiple Nobel lineages?” — these are graph traversal problems that benefit from a combination of traversal queries and graph algorithms.
Let’s begin by building this graph in Ladybug.
Preprocessing the Data#
The JSON is nested: each record contains a children array (mentees) and a parents array (mentors). We need to flatten this into two DataFrames: nodes and edges.
Polars handles this with explode (fan array elements into rows) and unnest (promote struct fields into columns):
import polars as pl
df = pl.read_json(path)
df = df.explode("children").unnest("children")
df = df.filter(pl.col("type") == "laureate")
df = df.explode("parents")
df = df.with_columns(
pl.col("parents").struct.field("name").alias("parent_name"),
pl.col("parents").struct.field("id").alias("parent_id"),
).drop("parents")From this flattened DataFrame, we build node and edge tables. Laureates carry their prize category and year; scholars get nulls for those fields. Edges are deduplicated (mentor_id, mentee_id) pairs.
Since Polars is Arrow-native and Ladybug ingests Arrow directly, the data moves from DataFrame to graph with zero-copy semantics.
Loading into Ladybug#
With our DataFrames ready, we create the graph schema and load the data. Because Ladybug is embedded, all you do is define a connection to a local file named nobel.lbug. This is where the graph is stored.
import ladybug as lb
db = lb.Database("nobel.lbug")
conn = lb.Connection(db)
conn.execute("""
CREATE NODE TABLE Scholar(
id STRING PRIMARY KEY,
name STRING,
prize STRING,
year INT64,
is_laureate BOOLEAN DEFAULT false
);
""")
conn.execute("CREATE REL TABLE MENTORED(FROM Scholar TO Scholar);")Loading uses Ladybug’s LOAD FROM clause, which scans Polars DataFrames directly, and the MERGE command then writes the data to the database:
conn.execute("""
LOAD FROM nodes
MERGE (s:Scholar {id: id})
SET s.name = name, s.prize = prize, s.year = year, s.is_laureate = is_laureate;
""")
conn.execute("""
LOAD FROM edges
MATCH (s1:Scholar {id: from_id}), (s2:Scholar {id: to_id})
MERGE (s1)-[:MENTORED]->(s2);
""")Edges after unnesting JSON: 1324 rows
Unique nodes: 1513, Unique edges: 1314
shape: (1, 1)
┌──────────────┐
│ num_scholars │
│ --- │
│ i64 │
╞══════════════╡
│ 1513 │
└──────────────┘
shape: (1, 1)
┌─────────────────┐
│ num_mentorships │
│ --- │
│ i64 │
╞═════════════════╡
│ 1314 │
└─────────────────┘The result: 1,513 nodes and 1,314 edges, persisted to a local graph database on-disk.
Preparing the Schema for Algorithm Results#
The goal is to compute graph algorithm metrics using Icebug and write the computed metrics for each node back to the database. Because Ladybug is strictly typed, we need to define these new columns in the schema before we can write to them:
conn.execute("ALTER TABLE Scholar ADD IF NOT EXISTS pagerank DOUBLE DEFAULT 0.0;")
conn.execute("ALTER TABLE Scholar ADD IF NOT EXISTS betweenness DOUBLE DEFAULT 0.0;")Like any other strictly typed database, this is a one-time schema migration. The algorithms can be re-run as needed, writing fresh scores into these columns at a later time.
Running Graph Algorithms with Icebug#
Icebug ⤴ is a fork of networkit ⤴, the high-performance C++ graph analytics library. The fork optimizes for columnar data, emphasizing zero-copy processing with Apache Arrow to match Ladybug’s storage format.
The Icebug PageRank benchmark ⤴ on a 115M-node Wikidata graph shows an 8x speed advantage over networkit and 29% less memory usage. The columnar bet pays off.
First, we query the edge list from Ladybug and build an integer ID mapping (Icebug requires integer node IDs):
edge_df = conn.execute("""
MATCH (a:Scholar)-[:MENTORED]->(b:Scholar)
RETURN a.id AS src, b.id AS dst;
""").get_as_pl()
id_to_int = {node_id: i for i, node_id in enumerate(node_df["id"].to_list())}
G = nk.Graph(n, weighted=False, directed=True)
for src, dst in zip(edge_df["src"].to_list(), edge_df["dst"].to_list()):
G.addEdge(id_to_int[src], id_to_int[dst])Then we run PageRank and Betweenness Centrality:
pr = nk.centrality.PageRank(G)
pr.run()
bc = nk.centrality.Betweenness(G, normalized=True)
bc.run()PageRank measures influence: nodes with many incoming edges from other influential nodes score higher. Betweenness measures brokerage: nodes that sit on many shortest paths between other nodes score higher. We normalize betweenness to a [0,1] interval for easier comparison.
Finally, we write the scores back to Ladybug:
results = pl.DataFrame({
"id": [int_to_id[i] for i in range(n)],
"pagerank": pr.scores(),
"betweenness": bc.scores(),
})
conn.execute("""
LOAD FROM results
MERGE (s:Scholar {id: id})
SET s.pagerank = pagerank, s.betweenness = betweenness;
""")The scores now live in the graph, queryable alongside the original node properties.
What the Graph Tells Us#
Now we can query the graph to see who stands out. Top 5 Physics laureates by PageRank:
MATCH (s:Scholar)
WHERE s.prize = 'Physics'
RETURN s.name AS name, s.pagerank AS pagerank
ORDER BY s.pagerank DESC
LIMIT 5;The names: Serge Haroche, Claude Cohen-Tannoudji, Eric Cornell, Robert Millikan, Martin Perl. These are scholars with deep mentorship lineages feeding into them.
For betweenness centrality, we also pull mentor and mentee counts to see why these nodes matter:
MATCH (a:Scholar)-[:MENTORED]->(s:Scholar)-[:MENTORED]->(b:Scholar)
WHERE s.prize = 'Physics'
RETURN s.name AS name,
s.betweenness AS betweenness,
COUNT(DISTINCT a) AS num_mentors,
COUNT(DISTINCT b) AS num_mentees
ORDER BY betweenness DESC
LIMIT 5;┌──────────────────┬─────────────┬─────────────┬─────────────┐
│ name ┆ betweenness ┆ num_mentors ┆ num_mentees │
│ --- ┆ --- ┆ --- ┆ --- │
│ str ┆ f64 ┆ i64 ┆ i64 │
╞══════════════════╪═════════════╪═════════════╪═════════════╡
│ Max Born ┆ 0.00016 ┆ 6 ┆ 8 │
│ Joseph Thomson ┆ 0.000152 ┆ 4 ┆ 11 │
│ Niels Bohr ┆ 0.000108 ┆ 3 ┆ 10 │
│ Isidor Rabi ┆ 0.000083 ┆ 4 ┆ 3 │
│ Julian Schwinger ┆ 0.000051 ┆ 2 ┆ 5 │
└──────────────────┴─────────────┴─────────────┴─────────────┘The top betweenness scorers: Max Born (6 mentors, 8 mentees), J.J. Thomson (4 mentors, 11 mentees), Niels Bohr (3 mentors, 10 mentees), Isidor Rabi, Julian Schwinger. These are the bridges of 20th century physics.
The Thomson → Bohr → Rabi chain is particularly striking: it connects early 20th century British physics through Copenhagen to the Manhattan Project era. Three nodes, three generations, spanning the birth of quantum mechanics to the atomic age.
The two metrics reveal different things. PageRank captures downstream influence: who spawned the biggest tree of academic descendants. Betweenness captures brokerage: who connected different clusters of researchers. The most influential mentors aren’t always the most central bridges, and vice versa.
You can explore the graph visually using Ladybug Explorer ⤴:
docker compose upOpen http://localhost:8000 ⤴ in your browser to interact with the Nobel laureate mentorship network. The explorer runs in read-only mode, so you can safely query and visualize the data without modifying it.
MATCH (a:Scholar)-[r:MENTORED*1..2]->(b:Scholar)
WHERE a.prize = 'Physics' AND b.prize = 'Physics'
AND a.name = "Joseph Thomson"
RETURN * LIMIT 100;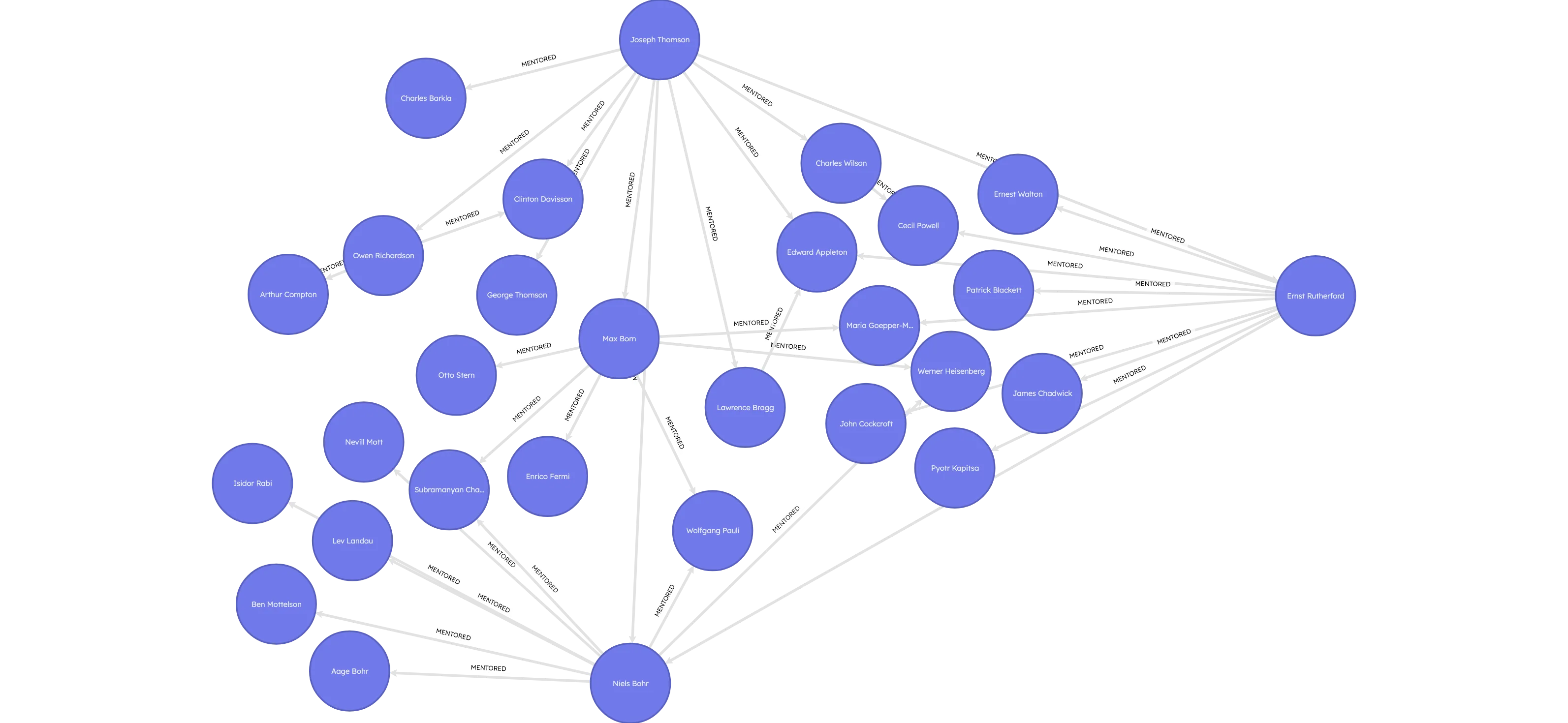
The Bigger Picture: Ladybug Memory#
What we’ve walked through is a small example: load JSON, build a graph, run algorithms, query results. But it demonstrates a pattern that Ladybug Memory is building toward at a much larger scale.
The problem Ladybug is solving is much larger: the integration between embedded relational databases like Postgres and DuckDB and the graph database ecosystem isn’t great. DuckDB-adjacent companies like MotherDuck focus on ease of deploying on the cloud and offering managed database solutions, not local-first storage. On the trajectory that Kuzu was once headed, it was aiming to offer much the same, but for graph database users.
However, we now live in an agentic era. The primary need of agent builders is to persist structured memory locally (or on the cloud, if required). They also need to query it efficiently, and run analytics without constantly moving data between formats and external systems. This leads to the vision of Ladybug Memory ⤴.
Ladybug Memory’s platform combines four components:
- Pgembed handles transactional writes from multiple agents
- DuckDB auto-migrates aging data to columnar format
- Icebug discovers patterns via zero-copy graph analytics
- LadybugDB stores and queries the graph
The vision blog ⤴ for Ladybug Memory outlines a unified memory substrate for AI agents. This goes beyond “Graph RAG” to offer a full persistence layer that combines probabilistic and deterministic structures for LLMs to infer from, with a columnar graph format underneath. The data representation stays consistent whether it’s on disk or in memory, and while the system is local-first, it remains compatible with object stores and cloud infrastructure when needed.
On the roadmap: SparseTensors for embedding storage, GPU acceleration for larger-scale analytics, and continued work toward graph-native AI infrastructure.
Get Involved#
Ladybug is open source. Whether you’re interested in graph algorithms, columnar storage, or building memory infrastructure for AI agents, there’s room to get involved.
To try what we walked through in this post, check out the code for this tutorial ⤴. For the broader ecosystem that Ladybug Memory builds on top of, check out these links:
- LadybugDB docs ⤴ and GitHub ⤴
- Icebug on GitHub ⤴
- Pgembed on GitHub ⤴
- Ladybug Memory platform ⤴ and vision blog ⤴
If you’re building agent memory, have heard about “context graphs”, or have opinions about what graph-native memory should look like, join the Ladybug community on Discord ⤴ to chat more!